Secondary metabolite discovery and a plant wall biodesign opportunity
May 25, 2023
Pitch 1: Laura Carroll: Machine learning approaches for microbial secondary metabolite discovery
Assistant Professor in the Department of Clinical Microbiology at Umeå University and a SciLifeLab and Wallenberg Data-Driven Life Science (DDLS) Fellow
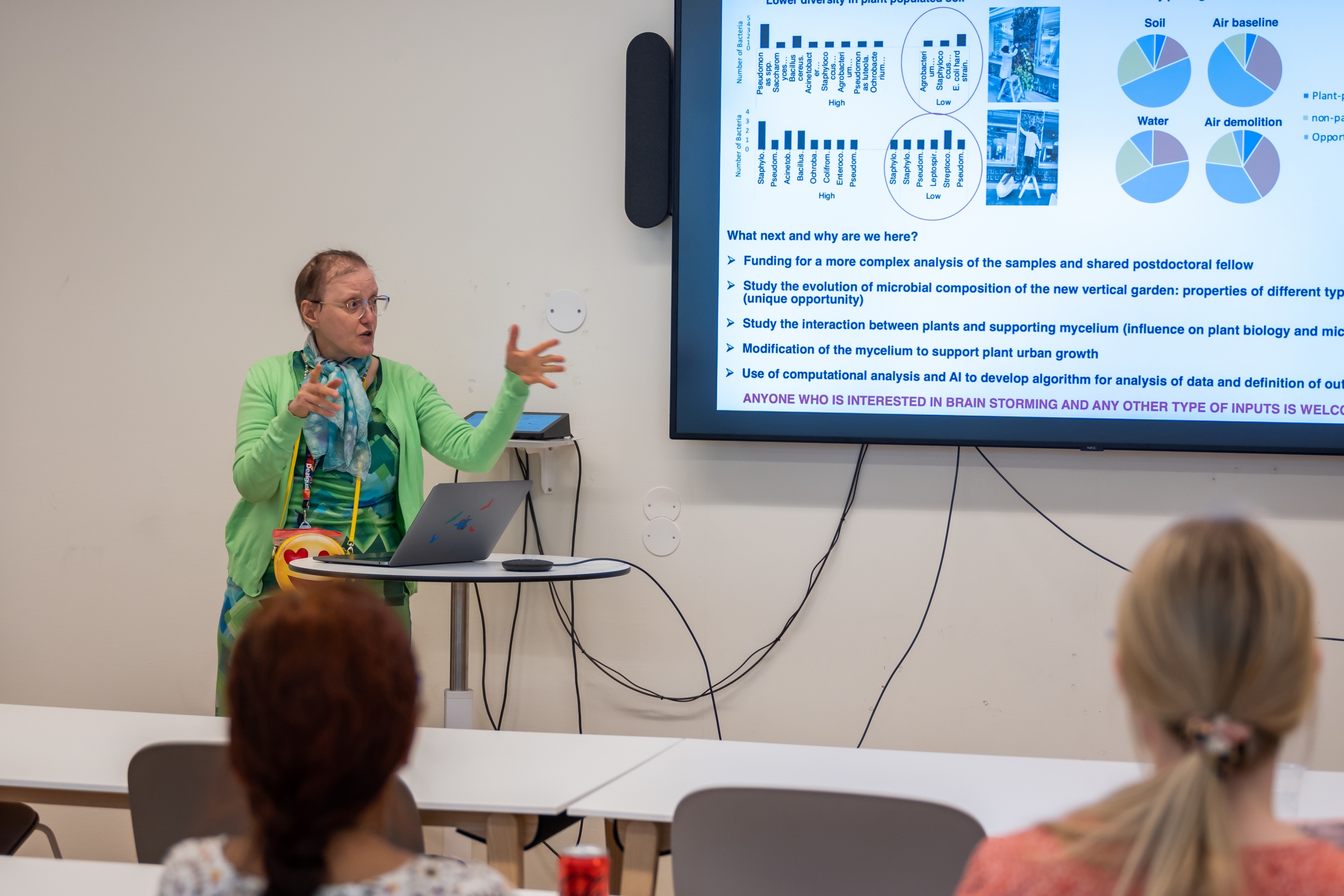
Pitch 2: Danielle Wilde and Teresa Frisan: Växtvägg – bringing micro and macro to the service of human-plant interactions
Danielle Wilde: Professor at Umeå Institute of Design (UID)
Teresa and Danielle co-lead collaborative research exploring how Molecular Biology and Design can collaborate effectively with other disciplines by facing a shared concern. They are currently working with SLU Forestry on a project that explores how to a) bring the boreal forest indoors, b) maximise food-based indoor plantings, and c) successfully grow other forms of vertical gardens in an indoor microclimate. This work is being developed at the UmArts campus at the site of a dying tropical plant wall.
The bacterial microenvironments of the dying wall are currently being characterised. Further samples will be taken as the new (demonstrator) wall begins to grow. This work aims to understand whether there is a correlation between plant survival and microbial microenvironment, which may be used to develop soil with a microbial composition that can support sustainable vertical gardens, culturally appropriate to Arctic settings.
Interested in: finding collaborators who can extend and deepen this work. One possibility is to undertake a microenvironmental analysis using artificial intelligence for identification of the sampled bacterial colonies. Other possibilities of extending the work are welcome!